Showcase
Page Info
Audience: Beginner
Prerequisites:
using PlantGeom, CairoMakieTime: 3 minutes
Output: Full-plant 3D visualizations from existing files
If you are new to programming, you can treat this page as "copy-paste and run" first. You can understand details later in the quickstarts.
Plant Visualization Showcase
A simple plant read from an OPF file and visualized with plantviz. The plant has explicit geometry in the file, so we can visualize it immediately without any additional steps. It is made of two internodes, each with one leaf:
using PlantGeom
using CairoMakie
files_dir = joinpath(dirname(dirname(pathof(PlantGeom))), "test", "files")
simple_opf = read_opf(joinpath(files_dir, "simple_plant.opf"))
plantviz(simple_opf, figure=(size=(980, 680),))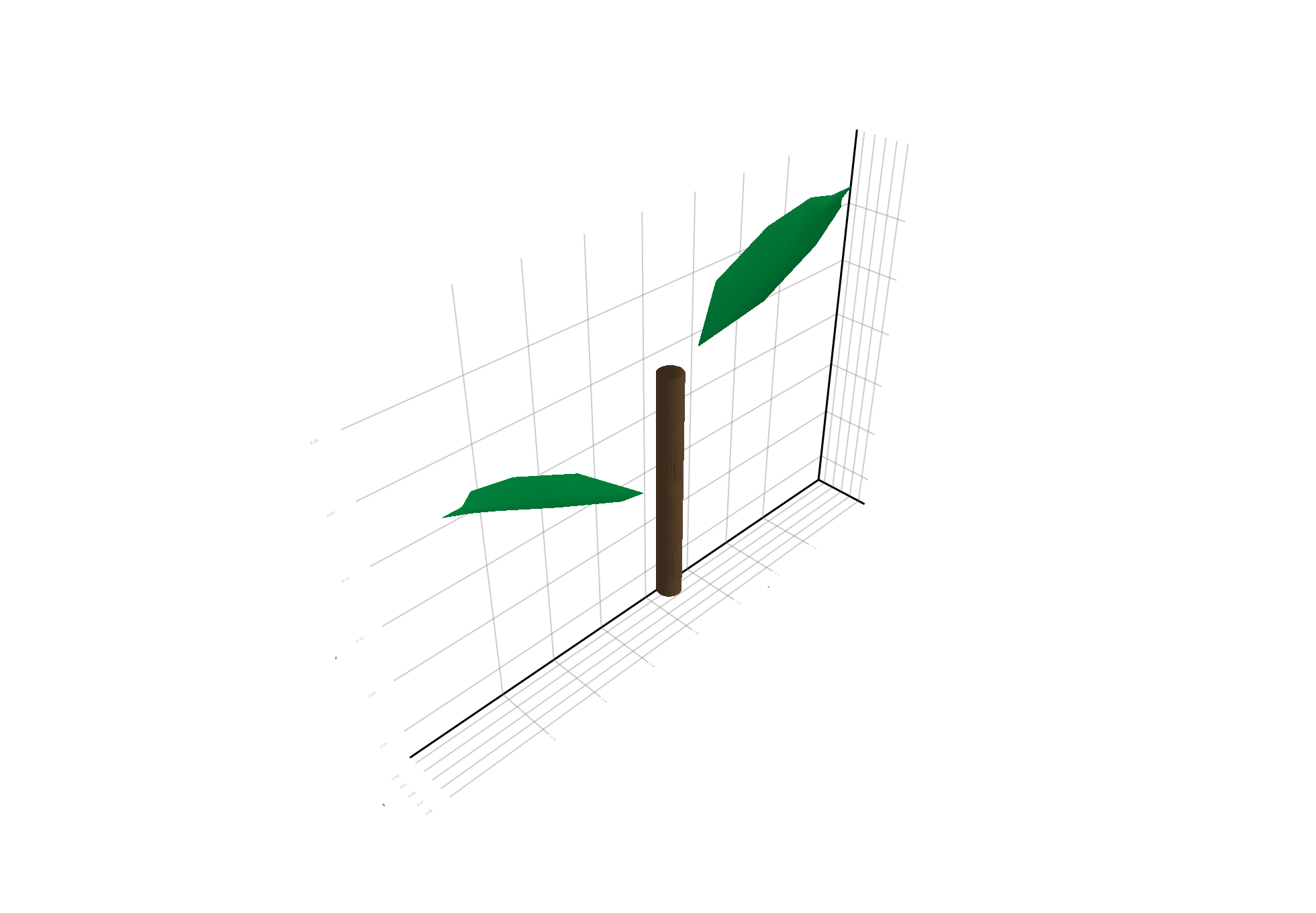
A more complex plant, also read from an OPF file. This plant is a coffee plant that was measured in the field and reconstructed:
coffee_opf = read_opf(joinpath(files_dir, "coffee.opf"))
plantviz(coffee_opf, figure=(size=(980, 720),))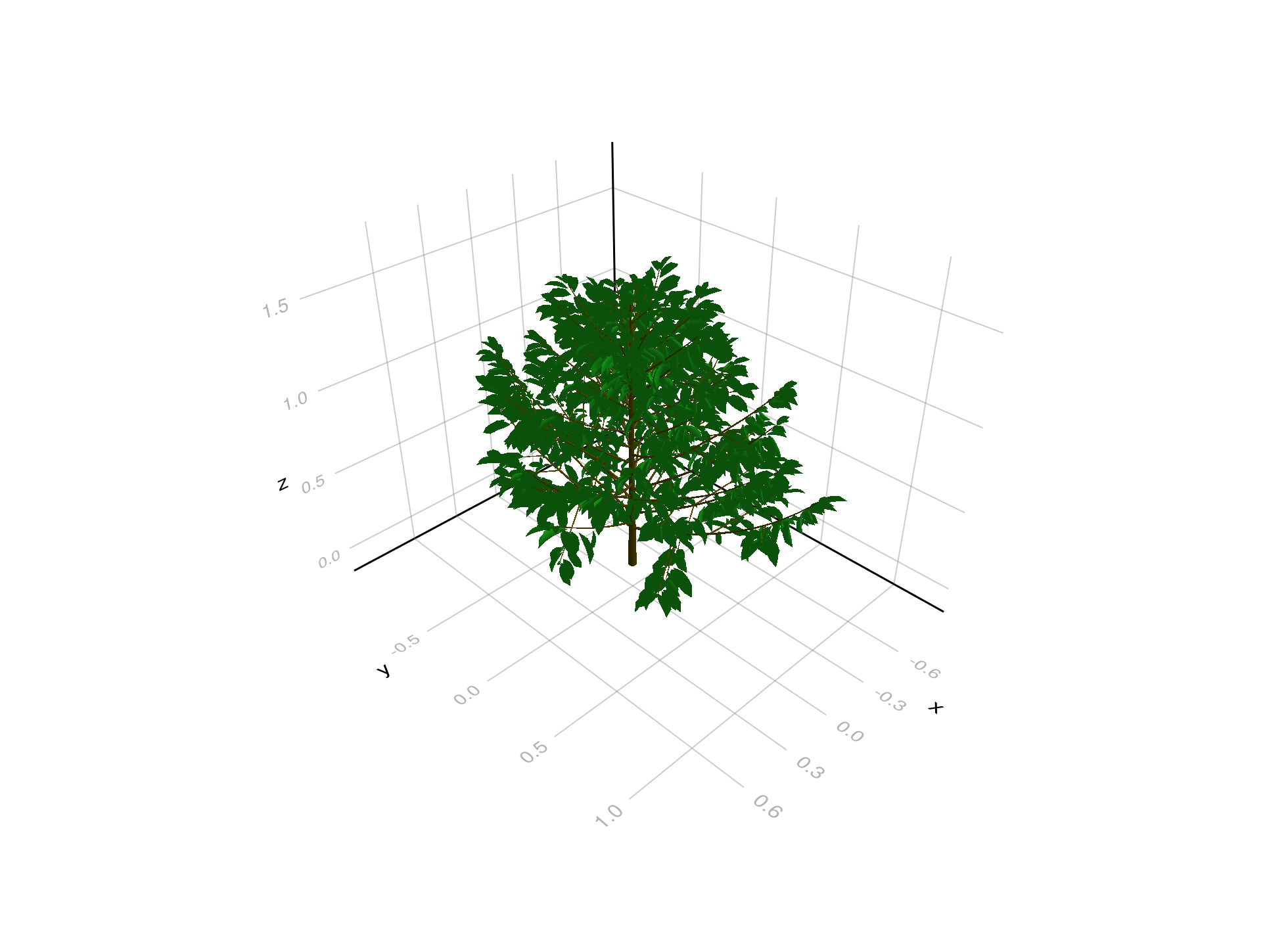
A tree generated with the growth API from PlantGeom:
tree_demo = build_demo_tree_with_growth_api()
plantviz(tree_demo, figure=(size=(980, 920),))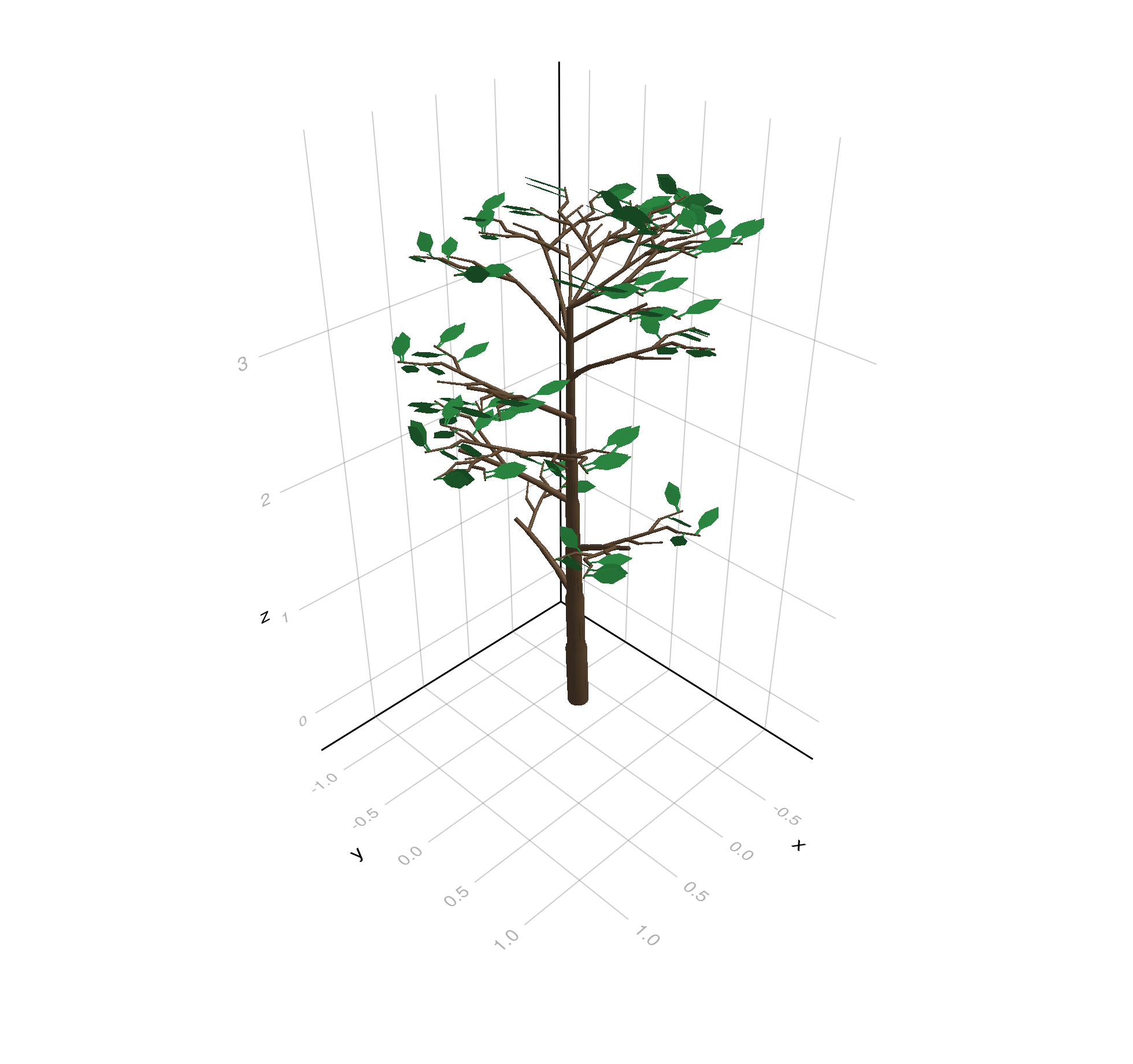
And an even more complex plant: a palm generated with the VPalm module from XPalm.jl, managed with PlantGeom geometry, and rendered with RayMakie by Simon Danisch:
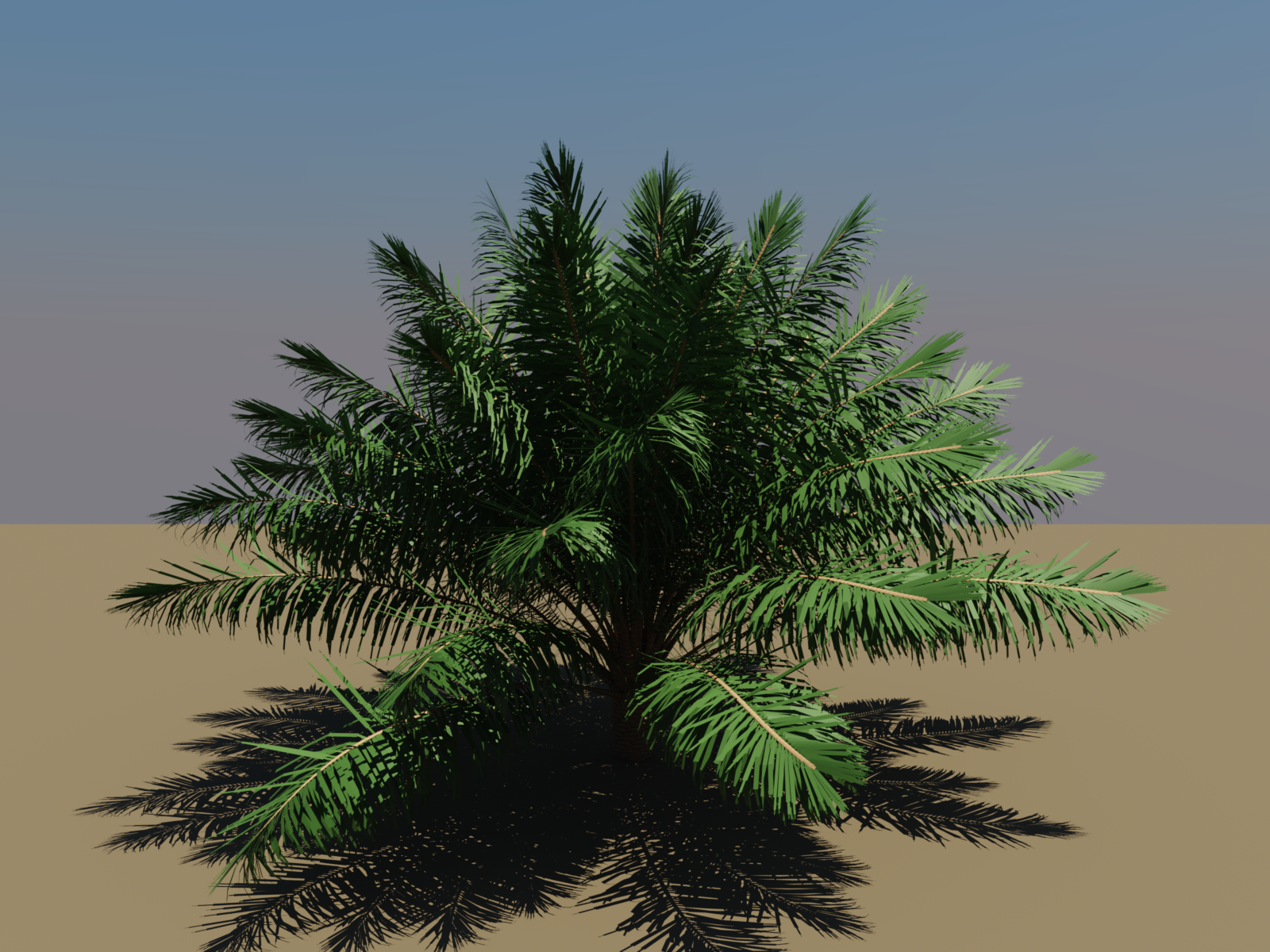

How It Works
read_opfgives you topology + geometry from existing plant files.build_demo_tree_with_growth_api()shows a generated plant using explicit growth + rebuild steps.plantvizmaterializes and renders current node geometries in one call.